highlights
Modes d’analyse de la variabilité des fréquences des polymorphismes génétiques dans les populations humaines
Skin colour patterns: predicting the unseen
In ‘Physical Review X (PRX)’, Milinkovitch's team predicts & confirms a secondary colour pattern in ocellated lizards. This pattern is too subtle for our eyes to see.
Understanding Architecture And Evolutionary Patterns In Haplolepidous Peristomes (Dicranidae, Bryophyta) Using Histology And Micro-Morphology
25.07.2023 14:00, Salle de conférence (Museum of Natural History)
Mathilde Ruche (Michelle Price's group).
hosted by: Michelle Price.
Research
Our department hosts 12 research laboratories gathering close to 200 scientists, engineers and technical staff. Research topics cover a large variety of topics, such as developmental genetics and neurogenetics, regeneration, evo-devo, physics of biology, phylogenetics or anthropology.
moreevents
-
25 Jul
Understanding Architecture And Evolutionary Patterns In Haplolepidous Peristomes (Dicranidae, Bryophyta) Using Histology And Micro-Morphology
-
30 Aug
to be announced
-
29 Sep
Mechanobiology of cell shape control
contact
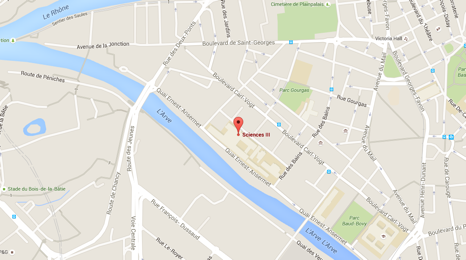
Department of Genetics and Evolution
Quai Ernest-Ansermet, 30
1205 Geneva
Switzerland
office: 4002A
T: +41 22 379 67 85
 more
more
