Keywords: hydra, cnidarian, regeneration, cellular remodelling, developmental plasticity, homeostasis, injury, wound healing, growth control, patterning, apoptosis-induced compensatory proliferation, signaling, evolution, neurogenesis
1. Summary
A fascinating, unanswered question in biology is how some organisms respond to injury by regenerating the missing body structure, whereas other have lost this potential. Here we propose to use the freshwater Hydra polyp as regeneration model organism to tackle this question. Indeed, Hydra is a simple animal that provides a powerful model system to understand how a highly dynamic homeostasis contributes to link wound healing to tissue repair and regeneration. The questions we address in our research are the following:
- HOMEOSTASIS & REGENERATION: What are the cellular and molecular mechanisms that maintain homeostasis in Hydra and drive regeneration after bisection?
- STEM CELLS: What are the respective functions of the stem cells and the differentiated cells in these processes?
- MEMORY: What are the memory mechanisms that allow the regenerating tip to regrow the appropriate missing structure, i.e. a head on one side and a foot on the other side of the bisection?
- ADULT NEUROGENESIS: What is the genetic circuitry that leads to de novo neurogenesis in cnidarians?
- AGING: What are the mechanisms that keep the developmental program(s) such as head regeneration, foot regeneration and asexual reproduction (budding) accessible all along the hydra life?
- EVOLUTION: Which of these mechanisms have been conserved along evolution?
To elucidate these questions, we want identify thanks to RNA interference (RNAi) the signaling cascades that govern cellular and developmental plasticity in Hydra. We recently showed that apoptosis-induced compensatory proliferation provides a mechanism to launch a complex regeneration program such as head regeneration. Interestingly apoptosis-induced compensatory proliferation seems to be also at work when Xenopus regenerates its tail, when Drosophila larvae regenerates their imaginal discs or even when rodents regenerate their skin or their liver (for recent reviews see [1, 2]). These results suggest that there might be some common paths to launch a regenerative response.
2. The Hydra model system in few words
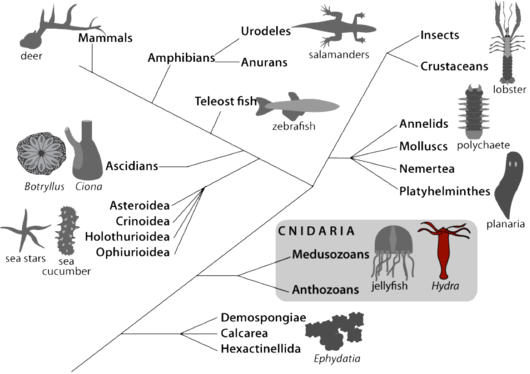
Hydra belongs to Cnidaria, a sister group to bilaterians (Fig.1). The anatomy of Hydra is rather simple: this is basically a bilayered tube that shows an apical-basal polarity, with at one extremity a head region that includes a mouth/anus opening and a ring of tentacles to actively catch the food (Hydra are carnivorous), and at the other extremity a basal disc. Its cellular organization is rather simple, with two cell layers, the ectoderm and the endoderm separated by an extracellular collagenoeous matrix, the mesogolea. Hydra differentiates all cell types required for neuro-muscular transmission, digestion, secretion and sexual reproduction. These different cell types differentiate from three distinct stem cell populations: myoepithelial cells located in the ectoderm, myoepithelial cells located in the endoderm and interstitial cells that are multipotent stem cells giving rise to neurons, mechano-sensory cells (nematocytes or cnidocytes), gland cells and gametes [3, 4].
Over the past 25 years, the genes encoding the signaling proteins that control and execute various cellular behaviors and developmental processes were shown to be present in all animals. The recent sequencing of the genome of two cnidarian species (Nematostella vectensis – sea anemone - and Hydra magnipapillata) indeed confirmed this high level of gene conservation from cnidarians to vertebrates. This definitively supports the relevance of Hydra as a model system to investigate complex biological questions [5, 6].
3. Functional analysis of the genetic circuitry supporting regeneration in Hydra
Gene silencing through RNAi by feeding identifies cellular phenotypes that are shared between Hydra and man
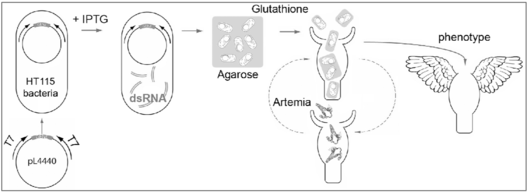
RNAi gene silencing obtained by feeding the animal on dsRNA-producing bacteria was first described in C. elegans and more recently adapted to planarians [7-9]. We adapted this strategy to Hydra (Fig. 2) and proved that this method that is harmless, stepwise and efficient, can induce gene-specific phenotypes [10-13]. In case of the protease inhibitor Kazal1that isspecifically expressed in gland cells of the gastrodermis, RNAi silencing induces a massive autophagy that extends to the digestive cells. This phenotype, which mimics the human SPINK1/mouse SPINK3 pancreatic phenotype, provides the first example of a conserved cellular mechanism from cnidarians to mammals, an example that definitely strengthen the paradigmatic value of this little animal.
This strategy offers the possibility of systematic RNAi screens in Hydra.
4. The apoptosis-dependent activation of the Wnt pathway in head-regenerating tips.
The canonical Wnt pathway is perfectly conserved in Hydra and required for the head to regenerate properly [14, 15]. We recently showed that an asymmetrical wave of apoptosis occurs immediately after mid-gastric bisection affecting 50% of the cells in head-regenerating tips but less than 7% in foot-regenerating ones [13]. Apoptotic cells actually transiently release the Wnt3 signal that activates b-catenin in the surrounding S-phase cells. Indeed progenitors migrate towards the wound, accumulate underneath the apoptotic zone and rapidly divide forming a proliferative zone. Upon inhibition of apoptosis by caspase inhibitors, or upon Wnt3 or b-catenin RNAi silencing, cell proliferation and head regeneration are abolished, whereas simply adding exogenous Wnt3 fully rescues these processes. Conversely
induction of apoptosis in foot-regenerating tips converts them to regenerate heads through the activation of the Wnt3/b-catenin pathway. Hence the level of apoptosis appears critical to trigger compensatory proliferation and head regeneration through the Wnt pathway.
The immediate asymmetric regulation of the MAPK ➜ RSK ➜ CREB ➜ CBP pathway
The MAPK/CREB pathway seems to play a key role in the induction of the head regeneration process. Indeed the cAMP Response Element Binding (CREB) protein is a transcription factor that likely interacts with distinct partners in CRE-binding complexes immediately after bisection [16], and exhibits a series of immediate regulations in the head- but not in the foot-regenerating tips as an RSK-dependent phosphorylation [17] and a rapid up-regulation of the CREB gene expression [18]. In vertebrates, the phosphorylated form of CREB binds to the chromatin modifyer CBP in order to modulate gene expression. Similarly the Hydra CBP encodes a CREB-binding domain and silencing of either RSK or CREB or CBP prevents the immediate wave of apoptosis and the cellular reorganization normally observed in head-regenerating tips (Chera and Galliot, submitted). We are currently investigating how this signaling pathway can sense the stress of amputation to activate a head regeneration program.
The emergence of neurogenesis and apical patterning in metazoans
In Hydra neurogenesis takes place continuously in the adult polyp but the dense apical nervous system also forms de novo in budding and regenerating animals [19- 21]. In bilaterian animals whose ancestor already had a centralized nervous system and sensory organs, the homeobox genes of the ANTP and PAIRED class play a major role in the developing central nervous system [22-24]. Each class includes a large number of families that for most of them diversified quite early in animal evolution, preceding the divergence of Cnidaria [25-27]. This raises the question of the function of these gene families in cnidarians (that differentiate a sophisticated nervous system) and in poriferans (that have no nervous system)[21].
The PAIRED-class homeobox genes
Among evolutionary conserved gene families involved in the specification of brain and sensory organs in bilaterians, several are expressed in the nervous system of cnidarians [28-32]. Among these, the paired-class homeobox gene prdl-a, is expressed in apical nerve cells in adult polyps but after bisection, transiently in endodermal cells of the head-regenerating tip. This result suggested that prdl-a is involved in Hydra head organizer activity through inductive interactions from endoderm to the overlying ectoderm [19, 28]. This result was puzzling as some paired- like genes perform similar tasks at the time head organizer activity is established in mammals, suggesting that molecular mechanisms of anterior patterning can be traced back to cnidarians [33].
The Hox/ParaHox genes
Most ANTP-class gene families do have cnidarian counterparts although Hox-related families are not all present and less similar. In fact Hox-like genes were not found in poriferan, therefore, cnidarian Hox-like genes can be considered as representative of the proto-Hox genes [26, 34-36]. Out of them, cnox-2, theortholog of the paraHox Gsx gene, stimulates interest as its expression is restricted to the precursors of the neuronal and nematocyte cell lineages and cnox-2 is also upregulated during de novo apical neurogenesis at the time head is forming, during budding and in regenerating polyps [11, 26, 37]. When cnox-2 expression is silenced via RNAi, alterations of the apical nervous system and apical patterning process are observed. In vertebrates and Drosophila, Gsx is also involved in neurogenesis [38]. These data suggest an evolutionarily-conserved function for cnox-2/Gsx in neurogenesis and a possible functional link between neurogenesis and apical patterning established quite early in animal evolution [11, 20, 36].
5. References
- Galliot, B., and Chera, S. (2010) The Hydra model: disclosing an apoptosis-driven generator of Wnt-based regeneration. Trends Cell Biol in press
- Galliot, B., and Ghila, L. (2010) Cell plasticity in homeostasis and regeneration. Mol Reprod Dev in press
- Steele, R.E. (2002) Developmental signaling in Hydra: what does it take to build a “simple” animal? Dev Biol 248, 199-219
- Galliot, B., Miljkovic-Licina, M., de Rosa, R., and Chera, S. (2006) Hydra, a niche for cell and developmental plasticity. Semin Cell Dev Biol 17, 492-502
- Putnam, N.H., Srivastava, M., Hellsten, U., Dirks, B., Chapman, J., Salamov, A., Terry, A., Shapiro, H., et al. (2007) Sea anemone genome reveals ancestral eumetazoan gene repertoire and genomic organization. Science 317, 86-94
- Chapman, J.A., Kirkness, E.F., Simakov, O., Hampson, S.E., Mitros, T., Weinmaier, T., Rattei, T., Balasubramanian, P.G., et al. (2010) The dynamic genome of Hydra. Nature 464, 592-596
- Timmons, L., Court, D.L., and Fire, A. (2001) Ingestion of bacterially expressed dsRNAs can produce specific and potent genetic interference in Caenorhabditis elegans. Gene 263, 103-112
- Kamath, R.S., Martinez-Campos, M., Zipperlen, P., Fraser, A.G., and Ahringer, J. (2001) Effectiveness of specific RNA-mediated interference through ingested double-stranded RNA in Caenorhabditis elegans. Genome Biol 2, RESEARCH0002
- Newmark, P.A., Reddien, P.W., Cebria, F., and Sanchez Alvarado, A. (2003) Ingestion of bacterially expressed double-stranded RNA inhibits gene expression in planarians. Proc Natl Acad Sci U S A 100 Suppl 1, 11861-11865
- Chera, S., de Rosa, R., Miljkovic-Licina, M., Dobretz, K., Ghila, L., Kaloulis, K., and Galliot, B. (2006)
Silencing of the hydra serine protease inhibitor Kazal1 gene mimics the human SPINK1 pancreatic phenotype. J Cell Sci 119, 846-857 - Miljkovic-Licina, M., Chera, S., Ghila, L., and Galliot, B. (2007) Head regeneration in wild-type hydra requires de novo neurogenesis. Development 134, 1191-1201
- Buzgariu, W., Chera, S., and Galliot, B. (2008) Methods to investigate autophagy during starvation and regeneration in hydra. Methods Enzymol 451, 409-437
- Chera, S., Ghila, L., Dobretz, K., Wenger, Y., Bauer, C., Buzgariu, W., Martinou, J.C., and Galliot, B. (2009) Apoptotic cells provide an unexpected source of Wnt3 signaling to drive hydra head regeneration. Dev Cell 17, 279-289
- Hobmayer, B., Rentzsch, F., Kuhn, K., Happel, C.M., von Laue, C.C., Snyder, P., Rothbacher, U., and Holstein, T.W. (2000) WNT signalling molecules act in axis formation in the diploblastic metazoan Hydra. Nature 407, 186-189
- Lengfeld, T., Watanabe, H., Simakov, O., Lindgens, D., Gee, L., Law, L., Schmidt, H.A., Ozbek, S., et al. (2009) Multiple Wnts are involved in Hydra organizer formation and regeneration. Dev Biol 330, 186-199
- Galliot, B., Welschof, M., Schuckert, O., Hoffmeister, S., and Schaller, H.C. (1995) The cAMP response element binding protein is involved in hydra regeneration. Development 121, 1205-1216
- Kaloulis, K., Chera, S., Hassel, M., Gauchat, D., and Galliot, B. (2004) Reactivation of developmental programs: the cAMP-response element-binding protein pathway is involved in hydra head regeneration. Proc Natl Acad Sci U S A 101, 2363-2368
- Chera, S., Kaloulis, K., and Galliot, B. (2007) The cAMP response element binding protein (CREB) as an integrative HUB selector in metazoans: clues from the hydra model system. Biosystems 87, 191-203
- Miljkovic-Licina, M., Gauchat, D., and Galliot, B. (2004) Neuronal evolution: analysis of regulatory genes in a first-evolved nervous system, the hydra nervous system. Biosystems 76, 75-87
- Galliot, B., Quiquand, M., Ghila, L., de Rosa, R., Miljkovic-Licina, M., and Chera, S. (2009) Origins of neurogenesis, a cnidarian view. Dev Biol 332, 2-24
- Galliot, B. (2010) A Key Innovation in Evolution, the Emergence of Neurogenesis: Cellular and Molecular Cues from Cnidarian Nervous Systems. In Key Transitions in Animal Evolution (Schierwater, B., and De Salle, R., eds), 127-161, Science Publishers & CRC Press
- Pichaud, F., and Desplan, C. (2002) Pax genes and eye organogenesis. Curr Opin Genet Dev 12, 430-434
- Hirth, F., Kammermeier, L., Frei, E., Walldorf, U., Noll, M., and Reichert, H. (2003) An urbilaterian origin of the tripartite brain: developmental genetic insights from Drosophila. Development 130, 2365-2373
- Denes, A.S., Jekely, G., Steinmetz, P.R., Raible, F., Snyman, H., Prud'homme, B., Ferrier, D.E., Balavoine, G., et al. (2007) Molecular architecture of annelid nerve cord supports common origin of nervous system centralization in bilateria. Cell 129, 277-288
- Galliot, B., de Vargas, C., and Miller, D. (1999) Evolution of homeobox genes: Q50 Paired-like genes founded the Paired class. Dev Genes Evol 209, 186-197
- Gauchat, D., Mazet, F., Berney, C., Schummer, M., Kreger, S., Pawlowski, J., and Galliot, B. (2000)
Evolution of Antp-class genes and differential expression of Hydra Hox/paraHox genes in anterior patterning. Proc Natl Acad Sci U S A 97, 4493-4498 - Ryan, J.F., Burton, P.M., Mazza, M.E., Kwong, G.K., Mullikin, J.C., and Finnerty, J.R. (2006) The cnidarian-bilaterian ancestor possessed at least 56 homeoboxes. Evidence from the starlet sea anemone, Nematostella vectensis. Genome Biol 7, R64
- Gauchat, D., Kreger, S., Holstein, T., and Galliot, B. (1998) prdl-a, a gene marker for hydra apical differentiation related to triploblastic paired-like head-specific genes. Development 125, 1637-1645
- Lindgens, D., Holstein, T.W., and Technau, U. (2004) Hyzic, the Hydra homolog of the zic/odd- paired gene, is involved in the early specification of the sensory nematocytes. Development 131, 191-201
- Gauchat, D., Escriva, H., Miljkovic-Licina, M., Chera, S., Langlois, M.C., Begue, A., Laudet, V., and Galliot, B. (2004) The orphan COUP-TF nuclear receptors are markers for neurogenesis from cnidarians to vertebrates. Dev Biol 275, 104-123
- Stierwald, M., Yanze, N., Bamert, R.P., Kammermeier, L., and Schmid, V. (2004) The Sine oculis/Six class family of homeobox genes in jellyfish with and without eyes: development and eye regeneration. Dev Biol 274, 70-81
- Marlow, H.Q., Srivastava, M., Matus, D.Q., Rokhsar, D., and Martindale, M.Q. (2009) Anatomy and development of the nervous system of Nematostella vectensis, an anthozoan cnidarian. Dev Neurobiol 69, 235-254
- Galliot, B., and Miller, D. (2000) Origin of anterior patterning. How old is our head? Trends Genet 16, 1-5
- Chourrout, D., Delsuc, F., Chourrout, P., Edvardsen, R.B., Rentzsch, F., Renfer, E., Jensen, M.F., Zhu, B., et al. (2006) Minimal ProtoHox cluster inferred from bilaterian and cnidarian Hox complements. Nature 442, 684-687
- Chiori, R., Jager, M., Denker, E., Wincker, P., Da Silva, C., Le Guyader, H., Manuel, M., and Queinnec, E. (2009) Are Hox genes ancestrally involved in axial patterning? Evidence from the hydrozoan Clytia hemisphaerica (Cnidaria). PLoS ONE 4, e4231
- Quiquand, M., Yanze, N., Schmich, J., Schmid, V., Galliot, B., and Piraino, S. (2009) More constraint on ParaHox than Hox gene families in early metazoan evolution. Dev Biol 328, 173-187
- Schummer, M., Scheurlen, I., Schaller, C., and Galliot, B. (1992) HOM/HOX homeobox genes are present in hydra (Chlorohydra viridissima) and are differentially expressed during regeneration. Embo J 11, 1815-1823
- Mieko Mizutani, C., and Bier, E. (2008) EvoDevo: the origins of BMP signalling in the neuroectoderm. Nat Rev Genet 9, 663-677
