Laboratoire de régénération et neurogénèse adulte
Mots-clés: hydre, cnidaire, régénération, remodelage cellulaire, plasticité du développement, homéostase, blessure, réparation tissulaire, contrôle de la croissance tissulaire, mise en place du schéma corporel, prolifération compensatrice induite par l’apoptose, voies de signalisation, évolution, neurogénèse
1. Résumé
Une question fascinante en biologie est celle posée par la variation du potentiel de régénération à l’intérieur du royaume animal: en effet si de nombreuses espèces sont douées de régénération, la plupart des mammifères ont vraisemblablement perdu ce potentiel. Dans notre laboratoire nous utilisons l’Hydre d’eau douce qui fournit un modèle puissant pour comprendre l’équilibre entre le maintien d’un état homéostatique très dynamique et l’activation de programmes de régénération. Les questions sur lesquelles nous nous penchons sont les suivantes:
- HOMEOSTASIE et REGENERATION: Quels sont les mécanismes cellulaires et moléculaires qui maintiennent l’homéostasie et quell est leur rôle dans la régénération?
- CELLULES SOUCHES et CELLULES DIFFERENCIEES: Quelles sont les contributions respectives des cellules souches et des cellules différenciées dans ces processus?
- MEMOIRE: Quels sont les mécanismes de mémoire qui permettent d’activer le bon programme au bon endroit, c’est à dire la régénération du pied d’un côté, la régénération de la tête de l’autre?
- NEUROGENESE ADULTE: Quelle est la circuitrie génétique qui conduit à la neurogénèse de novo puisque l’animal reconstitue complètement son système nerveux apical lors de la régénération de la tête?
- VIEILLISSEMENT: Quels sont les mécanismes de non-vieillissement qui permettent de maintenir accessibles les programmes de développement, régénération et bourgeonnement, quelque soit l’âge de l’animal?
- EVOLUTION: Lesquels de ces mécanismes ont été conservés au cours de l’évolution?
Afin d’élucider ces questions, nous cherchons à identifier par interférence à l’ARN (ARNi) les cascades de signalisation qui gouvernent la plasticité cellulaire et développementale de l’Hydre. Nous avons récemment montré que la prolifération compensatrice induite par l’apoptose joue un rôle important pour régénérer une tête après bisection au milieu de la colonne corporelle. De façon intéressante, la prolifération compensatrice induite par l’apoptose joue également un rôle important chez le tétard de Xénope qui régénère sa queue, chez la larve de Drosophile qui régénère ses disques imaginaux ou même chez les rongeurs qui régénèrent leur peau ou leur foie. Ces avancées suggèrent qu’il existe des voies communes chez les animaux pour activer un processus de régénération (voir revues récentes [1, 2]).
2. Le modèle de l’Hydre en quelques mots
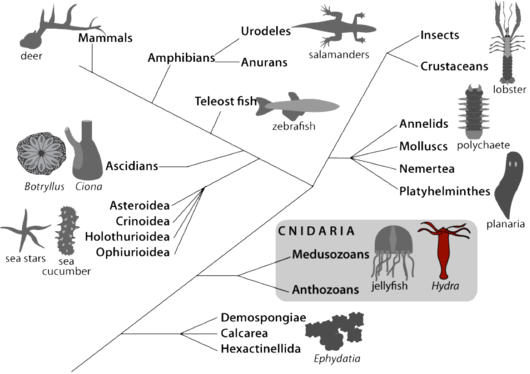
L’Hydre appartient au phylum des cnidaires, un groupe frère des bilatériens (Fig.1). L’anatomie de l’Hydre est simple: c’est en fait un tube qui est différencié aux deux extrémités. Au pôle apical l’animal différencie la région de la tête, formée d’un orifice bouche/anus entouré d’un cercle de tentacules qui servent à attraper la nourriture (l’Hydre est carnivore); au pôle basal, l’animal différencie un disque basal qui lui permet de se fixer. Les tissus sont formés de deux couches cellulaires, l’ectoderme et l’endoderme, séparées par une matrice extracellulaire la mésoglée. L’Hydre differencie tous les types cellulaires nécessaires à la digestion, la sécrétion, la transmission neuro-musculaire, la reproduction sexuée. Ces différents types cellulaires dérivent de trois populations dictinctes de cellules souches: les cellules épithéliales de l’ectoderme, les cellules épithéliales de l’endoderme et les cellules interstitielles qui sont multipotentes [3, 4]. En effet les cellules interstitielles peuvent tout au long de la vie de l’animal se différencier en cellules somatiques (neurones, cellules mécano-sensorielles - cellules urticantes nommées nématocytes ou cnidocytes -, cellules glandulaires) mais aussi en cellules germinales (ovocytes, spermatozoides).
Au cours des 25 dernières années, il a été montré que les gènes codant pour des protéines impliquées dans les voies de signalisation contrôlant les différents comportements cellulaires ou les processus de développement, avaient été extrêmement conservés au cours de l’évolution des métazoaires. Le séquençage du génome de deux espèces de cnidaires (Nematostella vectensis – anémone de mer - et tout récemment Hydra magnipapillata) a confirmé ce haut niveau de conservation génique entre les cnidaires et les vertébrés, renforçant encore la valeur du modèle de l’Hydre [5, 6].
3. Analyse fonctionnelle de la circuitrie génétique soutenant la régénération de l’Hydre
Grâce à l’ARN interférence (ARNi) par nourrissage nous avons pu mettre en évidence des phénotypes partagés par l’homme et l’Hydre
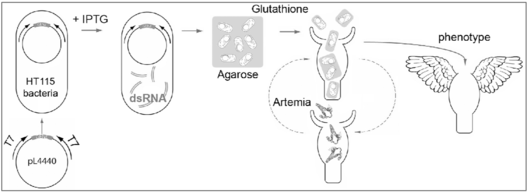
Chez certains vers, il est possible d’éteindre l’expression d’un gène donné en nourrissant les animaux avec des bactéries qui ont au préalable exprimé des ARN double-brins correspondant à ce gène [7-9]. Nous avons adapté cette stratégie à l’Hydre (Fig. 2) et ainsi pu démontrer qu’il s’agit d’une méthode non toxique pour l’Hydre, qui induit progressivement et efficacement des phénotypes spécifiques de chaque gène [10-13].
Dans le cas du gène Kazal1 qui produit un inhibiteur de protéases, l’ARNi induit une autophagie massive des cellules glandulaires qui produisent Kazal1 et des cellules digestives voisines qui normalement utilisent Kazal1. En effet en l’absence de Kazal1, les protéases ne sont pas bloquées et digèrent le contenu des cellules qui les produisent. Les souris mutées pour le gène de type Kazal SPINK3 développent également une autophagie massive au sein de leur pancréas et du tube digestif immédiatement après la naissance. Chez l’homme certaines pancréatites chroniques sont également dues à une autophagie excessive liée à des mutations du gène SPINK1. C’est la preuve de la conservation de la fonction des gènes de type Kazal de l’Hydre à l’homme, soutenant la valeur paradigmatique du système modèle de l’Hydre.
Cette stratégie de l’ARN interférence par nourrissage nous ouvre maintenant la possibilité de cribler un grand nombre de gènes pour identifier leur fonction au cours de la régénération.
L’Hydre utilise la mort cellulaire et l’activation de la voie Wnt3-bcatenin pour régénérer sa tête
L’activation de la cascade de signalisation Wnt3-bcatenin, qui est parfaitement conservée chez l’Hydre, est nécessaire à la régénération de la tête [14, 15]. Nous avons récemment montré que la bisection induit une vague d’apoptose dans les bourgeons destinés à régénérer une tête mais pas dans ceux destinés à régénérer un pied [13]. Ces cellules en apoptose jouent en fait un rôle moteur dans le processus de régénération car très transitoirement elles délivrent le signal Wnt3 aux cellules adjacentes qui se divisent alors rapidement. En effet si l’apoptose est bloquée ou si la production du signal Wnt3 est inhibée, la prolifération cellulaire et la régénération de la tête sont abolies mais l’addition de signal Wnt3 dans le milieu suffit à sauver le processus de régénération. Une autre expérience a mis en évidence cette fonction de la mort cellulaire dans le processus de régénération de la tête: il suffit d’induire l’apoptose dans les bourgeons destines à régénérer un pied pour activer la voie Wnt3-bcatenin et obtenir des hydres qui régénèrent une tête à la place du pied (Figure 3).
La régulation asymmétrique de la cascade MAPK ➜ RSK ➜ CREB ➜ CBP après bisection
La cascade MAPK/CREB semble jouer un rôle précoce dans l’induction de la régénération de la tête. En effet la protéine CREB (cAMP Response Element Binding protein) est un facteur de transcription qui interagit avec des partenaires différents lorsqu’il lie le motif CRE après bisection [16]. De plus CREB est soumis à une série de régulations différentes dans le bourgeon destiné à régénérer une tête et dans celui destine à régénérer un pied: du côté régénérant la tête CREB est immédiatement phosphorylé par la kinase RSK [17] et son expression est rapidement augmentée [18]. Chez les vertébrés, la forme phosphorylée de CREB lie le régulateur chromatinien CBP pour moduler l’expression de gènes cibles. La protéine CBP d’Hydre contient également un domaine liant CREB et lorsque l’expression des gènes RSK, CREB ou CBP est éteinte, ni la vague immédiate d’apoptose, ni le remodelage cellulaire normalement observé dans les bourgeons régénérant une tête n’ont lieu (Chera and Galliot, submitted). Nous cherchons actuellement à comprendre comment cette voie de signalisation permet de “lire” le stress de l’amputation pour activer un programme de régénération spécifique et complexe, celui de la formation de novo de la tête de l’Hydre.
4. L’émergence de la neurogénèse et de la différenciation apicale chez les métazoaires
Chez l’Hydre adulte la neurogénèse est constamment maintenue et le système nerveux apical très dense se forme au cours du bourgeonnement ou bien est complètement régénéré après décapitation [19-21]. Chez les animaux à symétrie bilatérale dont l’ancêtre commun avait déjà un système nerveux centralisé et des organes sensoriels, les gènes à homéoboite des classes ANTP et PAIRED jouent un rôle majeur au cours du développement du système nerveux [22-24]. Chaque classe comprend un grand nombre de familles qui, pour la plupart, se sont diversifiées très tôt au cours de l’évolution animale, précédant l’origine des cnidaires [25-27]. Ceci pose la question de la fonction de ces familles de gènes chez les cnidaires (qui différencient un système nerveux sophistiqué) et chez les porifères (qui n’ont pas de système nerveux) [21].
Les gènes à homéoboite de la classe PAIRED
Parmi les familles de gènes de la classe Paired impliquées dans la spécification du cerveau et des organes sensoriels chez les bilatériens, plusieurs sont exprimées dans le système nerveux des cnidaires [28-32]. Parmi ceux-ci, le gène prd-la est exprimé dans les précurseurs neuronaux et les neurones apicaux de l’Hydre adulte; cependant après bisection, ce gène est précocément et transitoirement exprimé dans la couche endodermale du bourgeon régénérant la tête. Cette double régulation suggère que ce gène est impliqué non seulement dans la neurogénèse mais également dans l’activité organisatrice de la tête en permettant des interactions inductives depuis l’endoderme vers l’ectoderme sus-jacent [19, 28]. Ce résultat est étonnant car chez les vertébrés, certains gènes paired-like ont une fonction similaire lorsque l’activité organisatrice de la tête se met en place au cours du développement précoce, suggérant des mécanismes moléculaires conservés [33].
Les gènes à homéoboite de la classe Hox/ParaHox
Si les cnidaires expriment un grand nombre de familles de gènes de la classe ANTP, toutes les familles Hox utilisées par les bilatéraux ne sont pas présentes et les gènes de type Hox n’ont pas été identifiés chez les porifères, suggérant que ces gènes “Hox-like” de cnidaires représentent en fait les gènes proto-Hox [26, 34-36]. Parmi ceux-ci, le gène cnox-2 (orthologue du gène ParaHox Gsx) est particulièrement intéressant car il est spécifiquement exprimé au cours de la neurogénèse apicale, chez le polype intact mais aussi au cours du bourgeonnement ou de la régénération de la tête [11, 26, 37]. Lorsque l’expression de cnox- 2 est éteinte par ARNi, le système nerveux apical n’est plus renouvellé et ne se forme plus correctement. Chez les vertébrés et chez la mouche Gsx est également impliqué dans la neurogénèse [38]. Ces données suggèrent une fonction neurogénique des gènes cnox- 2/Gsx très conservée au cours de l’évolution ainsi qu’un lien fonctionnel établi très tôt au cours de l’évolution animale entre neurogénèse et différenciation apicale/antérieure [11, 20, 36].
Références
- Galliot, B., and Chera, S. (2010) The Hydra model: disclosing an apoptosis-driven generator of Wnt-based regeneration. Trends Cell Biol in press
- Galliot, B., and Ghila, L. (2010) Cell plasticity in homeostasis and regeneration. Mol Reprod Dev in press
- Steele, R.E. (2002) Developmental signaling in Hydra: what does it take to build a “simple” animal? Dev Biol 248, 199-219
- Galliot, B., Miljkovic-Licina, M., de Rosa, R., and Chera, S. (2006) Hydra, a niche for cell and developmental plasticity. Semin Cell Dev Biol 17, 492-502
- Putnam, N.H., Srivastava, M., Hellsten, U., Dirks, B., Chapman, J., Salamov, A., Terry, A., Shapiro, H., et al. (2007) Sea anemone genome reveals ancestral eumetazoan gene repertoire and genomic organization. Science 317, 86-94
- Chapman, J.A., Kirkness, E.F., Simakov, O., Hampson, S.E., Mitros, T., Weinmaier, T., Rattei, T., Balasubramanian, P.G., et al. (2010) The dynamic genome of Hydra. Nature 464, 592-596
- Timmons, L., Court, D.L., and Fire, A. (2001) Ingestion of bacterially expressed dsRNAs can produce specific and potent genetic interference in Caenorhabditis elegans. Gene 263, 103-112
- Kamath, R.S., Martinez-Campos, M., Zipperlen, P., Fraser, A.G., and Ahringer, J. (2001) Effectiveness of specific RNA-mediated interference through ingested double-stranded RNA in Caenorhabditis elegans. Genome Biol 2, RESEARCH0002
- Newmark, P.A., Reddien, P.W., Cebria, F., and Sanchez Alvarado, A. (2003) Ingestion of bacterially expressed double-stranded RNA inhibits gene expression in planarians. Proc Natl Acad Sci U S A 100 Suppl 1, 11861-11865
- Chera, S., de Rosa, R., Miljkovic-Licina, M., Dobretz, K., Ghila, L., Kaloulis, K., and Galliot, B. (2006)
Silencing of the hydra serine protease inhibitor Kazal1 gene mimics the human SPINK1 pancreatic phenotype. J Cell Sci 119, 846-857 - Miljkovic-Licina, M., Chera, S., Ghila, L., and Galliot, B. (2007) Head regeneration in wild-type hydra requires de novo neurogenesis. Development 134, 1191-1201
- Buzgariu, W., Chera, S., and Galliot, B. (2008) Methods to investigate autophagy during starvation and regeneration in hydra. Methods Enzymol 451, 409-437
- Chera, S., Ghila, L., Dobretz, K., Wenger, Y., Bauer, C., Buzgariu, W., Martinou, J.C., and Galliot, B. (2009) Apoptotic cells provide an unexpected source of Wnt3 signaling to drive hydra head regeneration. Dev Cell 17, 279-289
- Hobmayer, B., Rentzsch, F., Kuhn, K., Happel, C.M., von Laue, C.C., Snyder, P., Rothbacher, U., and Holstein, T.W. (2000) WNT signalling molecules act in axis formation in the diploblastic metazoan Hydra. Nature 407, 186-189
- Lengfeld, T., Watanabe, H., Simakov, O., Lindgens, D., Gee, L., Law, L., Schmidt, H.A., Ozbek, S., et al. (2009) Multiple Wnts are involved in Hydra organizer formation and regeneration. Dev Biol 330, 186-199
- Galliot, B., Welschof, M., Schuckert, O., Hoffmeister, S., and Schaller, H.C. (1995) The cAMP response element binding protein is involved in hydra regeneration. Development 121, 1205-1216
- Kaloulis, K., Chera, S., Hassel, M., Gauchat, D., and Galliot, B. (2004) Reactivation of developmental programs: the cAMP-response element-binding protein pathway is involved in hydra head regeneration. Proc Natl Acad Sci U S A 101, 2363-2368
- Chera, S., Kaloulis, K., and Galliot, B. (2007) The cAMP response element binding protein (CREB) as an integrative HUB selector in metazoans: clues from the hydra model system. Biosystems 87, 191-203
- Miljkovic-Licina, M., Gauchat, D., and Galliot, B. (2004) Neuronal evolution: analysis of regulatory genes in a first-evolved nervous system, the hydra nervous system. Biosystems 76, 75-87
- Galliot, B., Quiquand, M., Ghila, L., de Rosa, R., Miljkovic-Licina, M., and Chera, S. (2009) Origins of neurogenesis, a cnidarian view. Dev Biol 332, 2-24
- Galliot, B. (2010) A Key Innovation in Evolution, the Emergence of Neurogenesis: Cellular and Molecular Cues from Cnidarian Nervous Systems. In Key Transitions in Animal Evolution (Schierwater, B., and De Salle, R., eds), 127-161, Science Publishers & CRC Press
- Pichaud, F., and Desplan, C. (2002) Pax genes and eye organogenesis. Curr Opin Genet Dev 12, 430-434
- Hirth, F., Kammermeier, L., Frei, E., Walldorf, U., Noll, M., and Reichert, H. (2003) An urbilaterian origin of the tripartite brain: developmental genetic insights from Drosophila. Development 130, 2365-2373
- Denes, A.S., Jekely, G., Steinmetz, P.R., Raible, F., Snyman, H., Prud'homme, B., Ferrier, D.E., Balavoine, G., et al. (2007) Molecular architecture of annelid nerve cord supports common origin of nervous system centralization in bilateria. Cell 129, 277-288
- Galliot, B., de Vargas, C., and Miller, D. (1999) Evolution of homeobox genes: Q50 Paired-like genes founded the Paired class. Dev Genes Evol 209, 186-197
- Gauchat, D., Mazet, F., Berney, C., Schummer, M., Kreger, S., Pawlowski, J., and Galliot, B. (2000)
Evolution of Antp-class genes and differential expression of Hydra Hox/paraHox genes in anterior patterning. Proc Natl Acad Sci U S A 97, 4493-4498 - Ryan, J.F., Burton, P.M., Mazza, M.E., Kwong, G.K., Mullikin, J.C., and Finnerty, J.R. (2006) The cnidarian-bilaterian ancestor possessed at least 56 homeoboxes. Evidence from the starlet sea anemone, Nematostella vectensis. Genome Biol 7, R64
- Gauchat, D., Kreger, S., Holstein, T., and Galliot, B. (1998) prdl-a, a gene marker for hydra apical differentiation related to triploblastic paired-like head-specific genes. Development 125, 1637-1645
- Lindgens, D., Holstein, T.W., and Technau, U. (2004) Hyzic, the Hydra homolog of the zic/odd- paired gene, is involved in the early specification of the sensory nematocytes. Development 131, 191-201
- Gauchat, D., Escriva, H., Miljkovic-Licina, M., Chera, S., Langlois, M.C., Begue, A., Laudet, V., and Galliot, B. (2004) The orphan COUP-TF nuclear receptors are markers for neurogenesis from cnidarians to vertebrates. Dev Biol 275, 104-123
- Stierwald, M., Yanze, N., Bamert, R.P., Kammermeier, L., and Schmid, V. (2004) The Sine oculis/Six class family of homeobox genes in jellyfish with and without eyes: development and eye regeneration. Dev Biol 274, 70-81
- Marlow, H.Q., Srivastava, M., Matus, D.Q., Rokhsar, D., and Martindale, M.Q. (2009) Anatomy and development of the nervous system of Nematostella vectensis, an anthozoan cnidarian. Dev Neurobiol 69, 235-254
- Galliot, B., and Miller, D. (2000) Origin of anterior patterning. How old is our head? Trends Genet 16, 1-5
- Chourrout, D., Delsuc, F., Chourrout, P., Edvardsen, R.B., Rentzsch, F., Renfer, E., Jensen, M.F., Zhu, B., et al. (2006) Minimal ProtoHox cluster inferred from bilaterian and cnidarian Hox complements. Nature 442, 684-687
- Chiori, R., Jager, M., Denker, E., Wincker, P., Da Silva, C., Le Guyader, H., Manuel, M., and Queinnec, E. (2009) Are Hox genes ancestrally involved in axial patterning? Evidence from the hydrozoan Clytia hemisphaerica (Cnidaria). PLoS ONE 4, e4231
- Quiquand, M., Yanze, N., Schmich, J., Schmid, V., Galliot, B., and Piraino, S. (2009) More constraint on ParaHox than Hox gene families in early metazoan evolution. Dev Biol 328, 173-187
- Schummer, M., Scheurlen, I., Schaller, C., and Galliot, B. (1992) HOM/HOX homeobox genes are present in hydra (Chlorohydra viridissima) and are differentially expressed during regeneration. Embo J 11, 1815-1823
- Mieko Mizutani, C., and Bier, E. (2008) EvoDevo: the origins of BMP signalling in the neuroectoderm. Nat Rev Genet 9, 663-677

